15 Open Source and Free Bioinformatics Tools List for Genomic Testing

India started genomic surveillance early this year to study the mutating nature of COVID-19. The surveillance strategy is aimed at performing genome sequencing as a quick response measure. And bioinformatics tools have come to the rescue here founder taking such genomic test.
What is Genomic Testing?
Genomic testing is performed to study mutations in a gene. Mutations indicate the presence of disorders and diseases, sometimes as deadly as cancer. Genomic sequencing tests identify differing levels of expression in a group of genes to understand their interactions. Test genomic analysis is thus critical for understanding the activity of genes and making an accurate prognosis.
What are the Different Types of Genomic Testing?
Genomic tests are a kind of medical tests done to rule out the possibility of a person having or developing a medical condition. These types of genomic testing are performed by geneticist or genetic counsellors:
- Diagnostic testing
Diagnostic testing is used for identifying genetic conditions that an individual may have. These are based on clinical presentations prepared by clinicians to confirm an initial diagnosis.
This kind of genomic test is done by checking an allele or a specific gene variant associated with a particular kind of disease. Bioinformatics tools here help with sequencing the genomes for further analysis.
- Clinical predictive testing
Clinical predictive kind of genomic testing is done to see if or not an individual is susceptible to a certain disorder. Open source and free bioinformatics tools here help with examining the causative gene variant for targeted tests.
- Pharmacogenomic testing
Open source and free bioinformatics tools for Linux help with pharmacogenomic testing by studying the genomic determinants of different drug responses. This test genomic analysis helps assess whether a particular medicine would be effective.
- Tumour testing
Tumour testing helps with sequencing of DNAs to study the mutations within them. Bioinformatics tools are deployed here to do tumour testing. This type of genomic testing is quite important when it comes to diagnosing cancers and tumours.
Advances in Genomic Testing to Analyse Health Risks
Genome-wide analysis and testing is critical for testing, monitoring and preventing diseases across populations.
Next generation genome sequencing technology undertakes parallel sequencing of DNA fragments for an efficient genomic analysis. These tests are performed using samples of hair, blood, tissue, amniotic fluid or skin. Some of the most advanced methods today for genomic testing are:
- Molecular tests for studying short length DNAs or single genes to examine mutations that may lead to genetic diseases.
- Biochemical test studies the activity level or number of proteins. Any abnormality here indicates the possibility of a genetic disorder.
Genomic testing has evolved newer advanced methods for analyzing the different health risks in new born. This is one area where genomic testing and medicine has advanced well and let us have a look here how:
- Presymptomatic & predictive testing
Presymptomatic & predictive testing is used in conditions where members of a family suffer from any kind of genetic disorder. This kind of genomic testing studies gene mutations to identify defects that may emerge either after birth or during later stages in life.
One good example could be identifying certain types of cancers using presymptomatic & predictive type of genomic testing.
- Carrier testing
People with increased risk of suffering from a genetic disorder are administered this type of genomic testing. Carrier testing assesses people who have more than one copy of genetic mutation.
- Prenatal testing
Prenatal kind of test screens a foetus for changes in her or his chromosomes/genes. The process makes it possible to ensure whether the child would have any genetic disorder.
- Chromosomal tests
Chromosomal tests analyze long length DNAs and whole chromosomes to find out if there are massive genetic changes.
How Bioinformatics Tools Help with Genomic Testing
Bioinformatics software solutions are ideal for genomic testing or next-generation sequencing. Next-generation sequencing technology is used to study mutations in genes for predicting the nature of a disease.
If we speak about this technology in the Indian context, you would find that India has started genome sequencing of COVID-19 using bioinformatics tools. Covid-19 has protein binding on its membrane and is primarily an RNA virus. It is because of these components that the virus has acquired the character it has and is fast mutating.
How biometric tools work is theydo genome sequencingto study the arrangement of DNA, RNA, and proteins in the virus to understand its changing nature. Bioinformatics tools with their next generation sequencing and genomic testing capabilities have been deployed by scientists in India thus to study such mutations.
Popular bioinformatics software solutions provide computation infrastructure for genomic data analysis.Bioinformatics tools are thus based on multiarray technology for transforming complicated genomic data into valuable insights.
Paid and open-source bioinformatics tools further support the sequencing data analysis for aligning and assembling DNA fragments. You can do variant calling, data visualizations, RNA expression profiling, gene fusion detections and data mining.
Bioinformatics tools help in the following fields for performing genomic tests:
Molecular biology- Bioinformatics software offers gRNA design tools for studying:
- Restriction enzyme map analysis
- Codeon frequency tables
- Sequencing primer design
- Sequence scrambler
Peptide- Bioinformatics tool offer peptide library design tools for:
- Amino acid code
- Peptide property calculator
- Antigen prediction
Common tools- Common tools in bioinformatics platform for Linux help with:
- Nucleic acid conversions
- Antibody drug discovery
Protein analysis- The application helps with protein and RNA analysis for:
- Psort II
- WOLF-PSORT
Bioinformatics Tools and Applications
Bioinformatics tools with their genomic testing abilities have been helpful in finding genetic alternations that have strong link to serious disorders and diseases. Test genomic tools has various applications:
- Sequence analysis- Sequence analysis is the method of subjecting an RNA, peptide sequence and DNA to different kinds of analytical methods. This is done to identify the origin, evolution and structure of biological databases.
- Molecular modelling– Molecular modelling makes use of computational and theoretical methodologies for analyzing the behaviour of molecules. Best open source and free bioinformatics tools use the simulation technique for performing molecular modelling.
- Molecular dynamics– Molecular dynamics application in bioinformatics determine the physical motion of atoms. The method is important for cell like environment in the field of structural bioinformatics. Measurements used here include graph theories, dynamic ross relations and perturbation response scans.
- FASTA in bioinformatics– FASTA is a format that is text-based in nature and is used to represent peptide and nucleotide sequences. FASTA software packages for bioinformatics tools help here with sequencing protein and DNA alignments.
- Phylogenetic analysis in bioinformatics– Phylogenetic analysis by bioinformatics tools provides branch diagrams for representing the relationship or evolutionary history between different organisms/species. These branch diagrams are called phylogenetic trees and they help identify characteristics like genes, organs, and proteins in an organization.
- Biological databases in Bioinformatics– Biological databases in bioinformatics can be best understood by analysing its three classifications that exist. The three classifications are- functional, structure and sequence. Protein sequences and nucleic acid is stored in sequence database whereas proteins & RNA exists in structural databases. Physiological role for gene products is provided by functional databases.
15 Free Bioinformatics Tools List 2026
Popular applications for bioinformatics are best for sequence analysis and curations. The best solutions in the field have key inbuilt computational and big data analysis tools for genome sequencing. Let us have a look at what else these applications are comprised of in the following list.
- geWorkbench
- BioPerl
- UGENE Open Source Bioinformatics Tool Linux
- Biojava Bioinformatics Tool for Linux
- Biopython Test Genomic Software
- InterMine
- IGV Genomic Sequencing Tool
- GROMACS
- Taverna Workbench
- EMBOSS Bioinformatics Tool Linux
- Clustal Omega
- BLAST
- Bedtool
- Bioclipse Open Source Bioinformatics Tool
- Bioconductor
geWorkbench
Best for: Hierarchical clustering
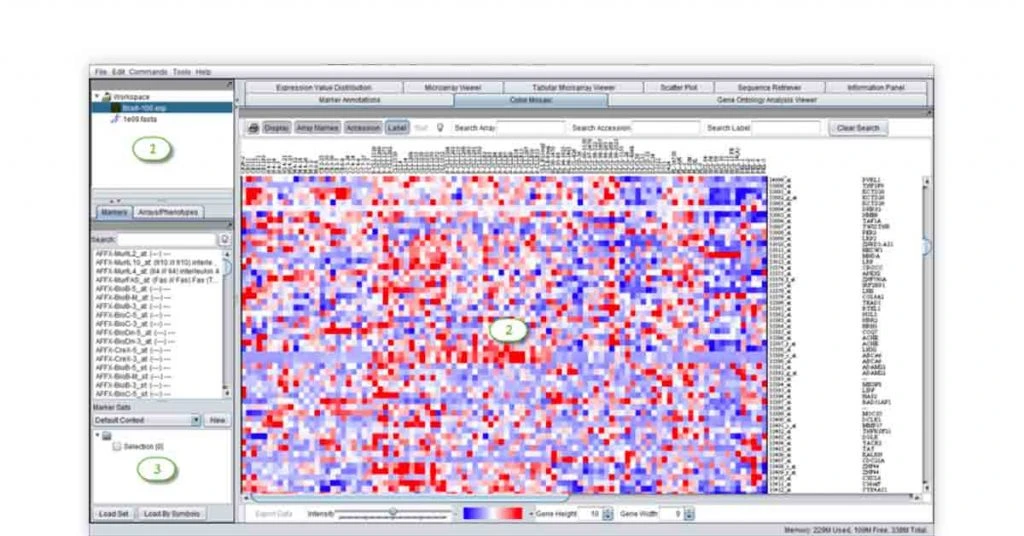
geWorkbench bioinformatics software offers computational and java-based tools for visualizing, supporting and analyzing sequence data. The bioinformatics tool also supports various plug-ins for genomics and gene integration.
The open-source and free bioinformatics tool can also be used for basic statistical analysis and transcription factor analysis.
Features of geWorkbench:
- Annotation pathways for collecting data
- Self-organizing maps & T-tests
- Hierarchical clustering
- Gene ontology enrichment analysis
- Microarray gene expression
- Network reverse engineering
geWorkbench Download: geWorkbench download available for Linux, Mac OS X and Windows.
BioPerl
Best for: Computational molecular biology
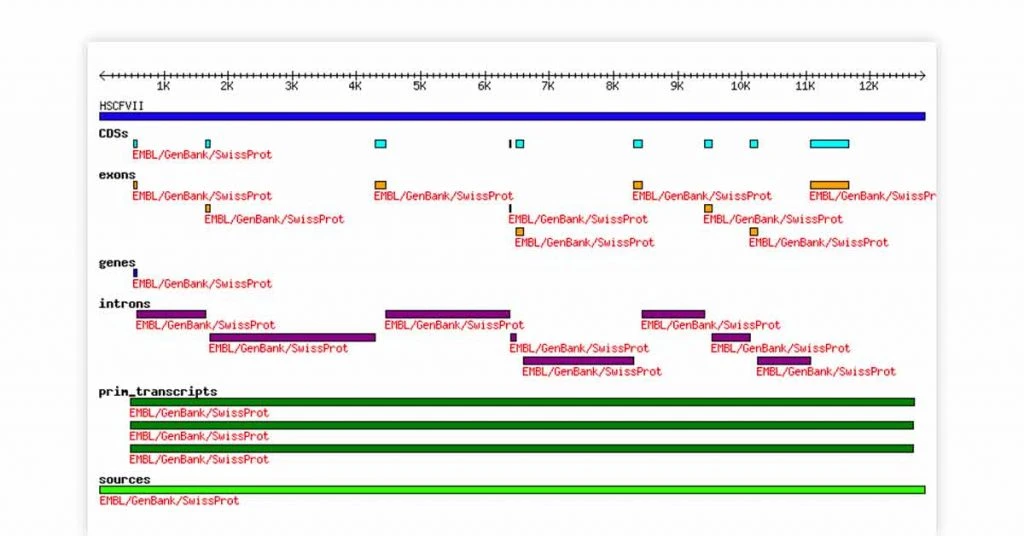
BioPerl bioinformatics tool for Linux is most deployed for computational molecular biology. The standardized CPAN style is the unique selling point of this bioinformatics platform. The Linux bioinformatics software offers Perl modules for peptide and nucleotide sequence data.
BioPerl Modules:
- Parsing real BLAST output
- Graphical rendering
- EST clusters
- Cytogenetic and radiation hybrid maps
BioPerl Download: You can install BioPerl on Linux, Unix and mac OS devices.
BioPerl Tutorial: Access BioPerl tutorial pdf.
UGENE Open Source Bioinformatics Tool for Linux
Best for: Dot plot & Chromatogram visualizations

UGENE open source and free bioinformatics tool Linux offers a common UI for integrating the platform with other bioinformatics applications. The software supports multiple formats for biological data available and you can retrieve it from remote locations. UGENE software sequence tool uses multicore GPUs and CPUs for the best performance.
Features of UGENE Open Source Bioinformatics Tool:
- Chromatogram visualization
- Multiple align editor
- Visual and interactive genomes
- 3D viewing in MMDB and PDB formats
- Phylogenetic tree view
- Dot plot visualization
- Query designer
UGENE Download: UGENE software free download available for Linux, macOS and Windows platforms.
Unipro UGENE Tutorial: Access Unipro UGENE tutorial here.
Biojava Bioinformatics Tool for Linux
Best for: Ensemble databases
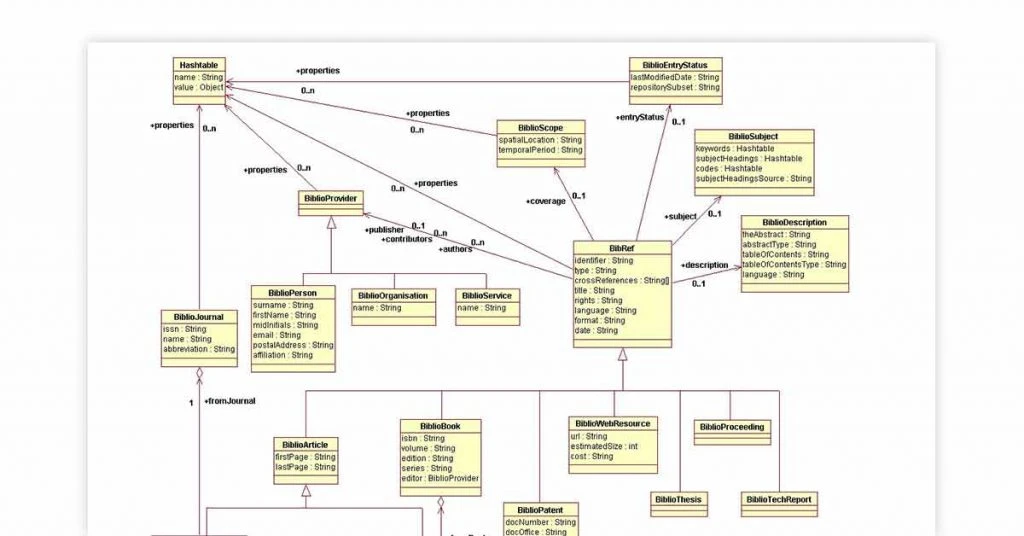
Biojava Bioinformatics tool for Linux, Windows and Solaris is best known for its various java tools ideal for processing biological data. The genome sequencing software supports a range of datasets and parsers for the common file format. Biojava test genomic tool can also be used for managing statistical and analytical routines.
Biojava Modules:
- Class objects and files for implementing java code
- Server support and DAS clients
- Ensembl databases
- Sequence alignments
- Homologene data
Updated Biojava Modules:
- Multiple structural alignments
- Support for protein assemblies
- Updated structure data model
- New structure file formats
- Accessible surface area
Biojava Download: Biojava downalod is available for Solaris, Windows and Linux.
Biojava Examples: Some good examples of Java library in Biojava are:
- Private static atom
- Public residue identifier
- Representative atom arrays
- String sequence
Biojava Tutorial: Go to GitHub’s link shared here for accessing Biojava tutorial.
Biopython Test Genomic Software
Best for: Performing sequence analysis in bioinformatics
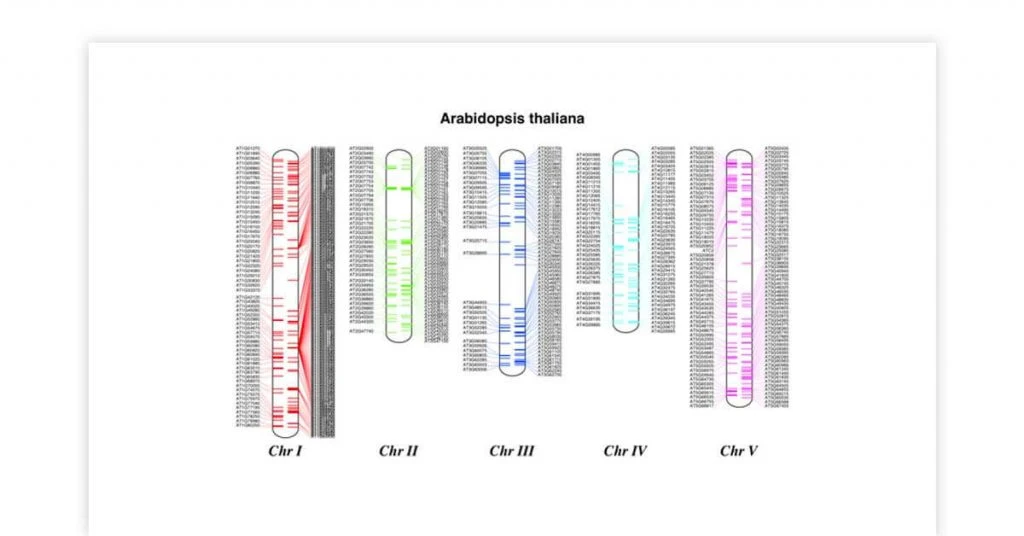
Biopython genome sequencing tool is most deployed for doing biological computation. This bioinformatics tool for Linux/UNIX supports multiple formats for bioinformatics files like FASTA, BLAST, Clustalw and Genbank. Inbuilt into Biopython are various python modules for making interactive and integrated sequences.
Features of Biopython Test Genomic Software:
- Managing sequence formats
- Protein structure
- Substitution matrices
- Weight calculations, transcriptions and translations
- EMBOSS command line tools
Biopython Uses:
- Microarray data for clustering
- Reading & writing tree-view files
- Structure data for PDB analysis and parsing
- BioSQL database
- Parser development & sequence plus options
Biopython GitHub: Biopython uses git as it its source code
Biopython Documentation: Clear documentation based on cookbook-style
Biopython Download: Download Biopython package for Unix/Linux platforms.
InterMine
Best for: Integrating biological data sources

InterMine bioinformatics tools is used for analyzing and integrating biological data. The open source and free bioinformatics platform supports dynamic tables for data filtering and drilling it down. The software functions by working with either a single object or multiple lists in multiple languages.
Features of InterMine:
- Search tools for templates, keywords and queries
- Support for GFF3, FASTA, Chado and such gene association files
- General python codes
InterMine GitHub: Create an account on GitHub accessing InterMine Python for python development.
InterMine Software Download: Youc can download InterMine by creating an account on GitHub.
InterMine Training Portal: InterMine training portal available for data query and analysis purposes.
IGV Genomic Sequencing Tool
Best for: Interactive genomics viewing
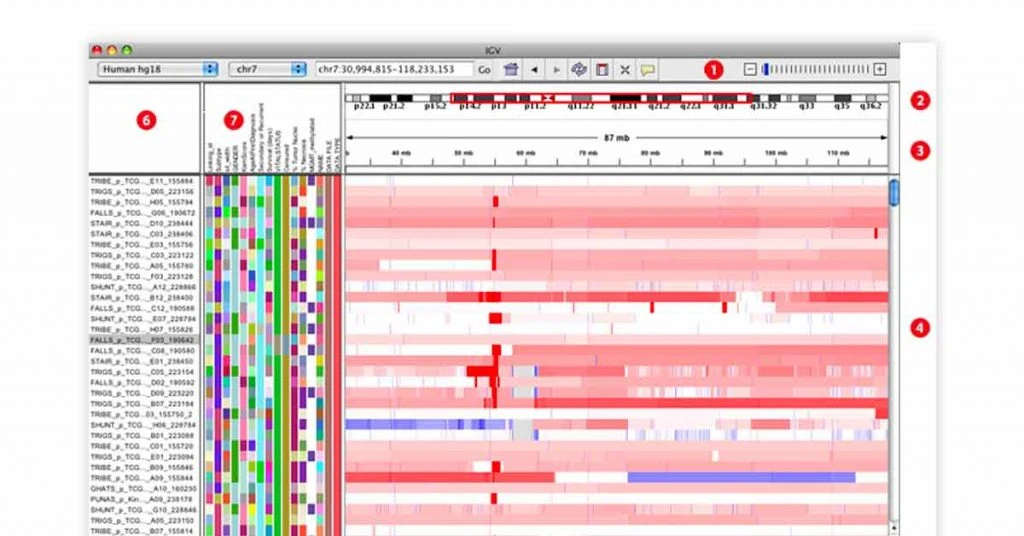
IGV genomic sequencing application offers genomics viewing for interactive genomics and effective visualization. IGV has been built on next-generation sequencing technology for genomic annotation. IGV bioinformatics software also lets you navigate through extensive data set and zoom it as per your requirement.
Features of IGV Test Genomic Tool:
- Attribute panel for tracking attributes
- Sorting and grouping tracked data
- File loading from genome space
- Flexible integration of genomic data/metadata
- Aligned sequence reads
- Data set loading from remote/local sources
- Multi-resolution file formats
IGV User Guide– Access IGV test genomic PDF.
GROMACS
Best for: Parameter files and topologies
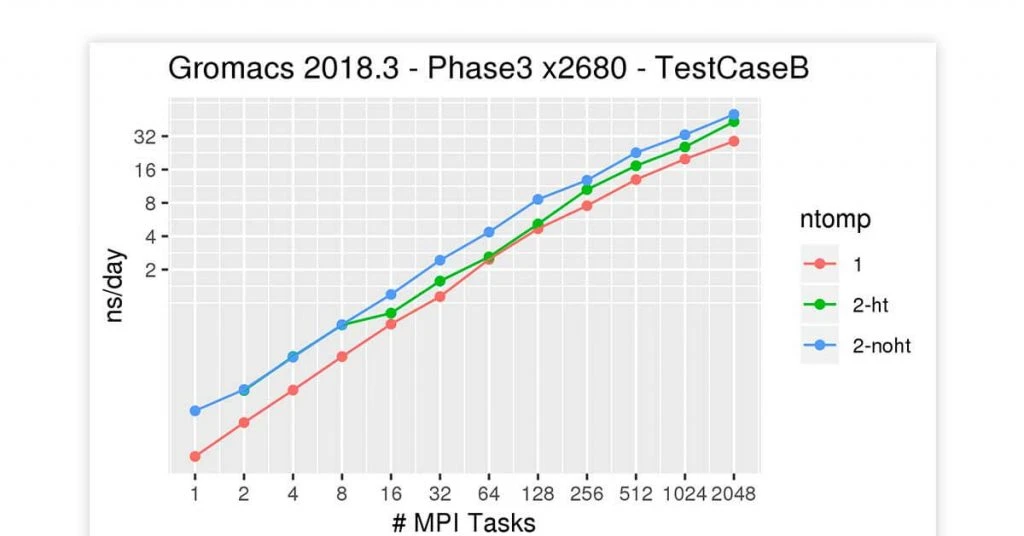
GROMACS bioinformatics tool for Linux offers building and analysis tools for dynamic molecular simulation. The bioinformatics software is used for molecular dynamics by simulating Newton’s motion from several particles. GROMACS performs well with lipids, proteins and such biochemical molecules for complex interactions.
Features of GROMACS:
- Parameter files and topologies
- File input & output through command line tools
- Trajectory and routine analysis
- MPI communication protocol
- State of the art algorithms
GROMACS Download: GROMACS download is available for Windows, macOS and Linux.
GROMACS Tutorial PDF: Access for GROMACS manual PDF.
Taverna Workbench
Best for: Silico experimentation
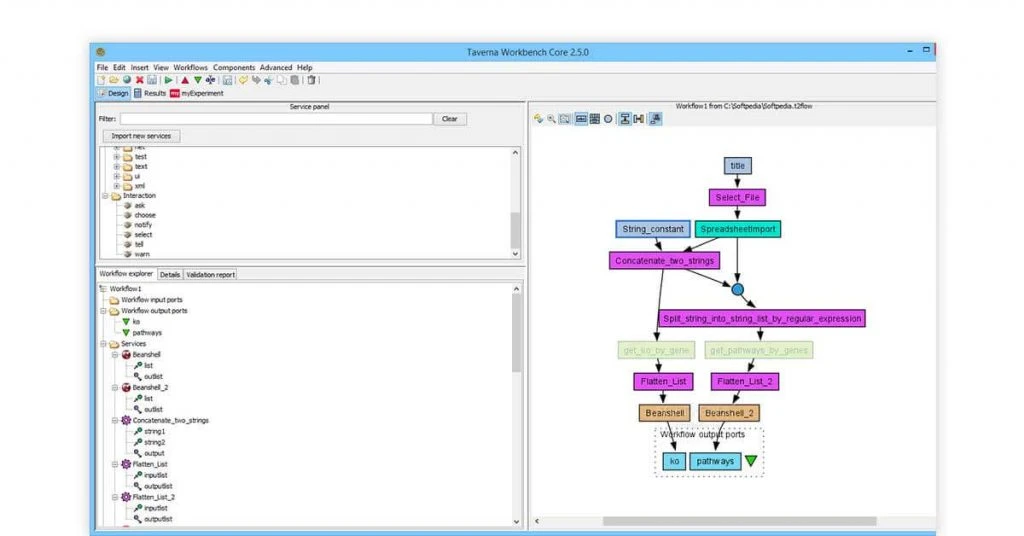
Taverna Workbench open source bioinformatics tool Linux is deployed for streamlining bioinformatics workflows. The best part about using Taverna Workbench is that it can be integrated with services like REST, WSDL and SOAP.
Taverna Workbench Features:
- Silico experimentation
- Command line executions
- Java libraries
- Semantically annotated components
- X.509 client authentication
- Debugging with workflow validation
Taverna Workbench Tutorial: Quick start guide for Taverna Workbench.
EMBOSS Bioinformatics Tool for Linux
Best for: Simplifying arrays and sequences
EMBOSS (European Molecular Biology Open Software Suite) bioinformatics platform is ideal for the field of molecular biology and sciences. EMBOSS is most used for web page data sequencing. You can also use this bioinformatics tools for nucleotide sequence pattern analysis.
Features of EMBOSS Open Source Bioinformatics Tool for Linux:
- Domain analysis
- Sequence alignment
- Protein motif identification
- Multiple structural and alignment formats
- Programming libraries for database indexing
- Simplified arrays and sequences
EMBOSS Software Download: Get EMBOSS software free download.
EMBOSS Bioinformatics Tutorial: Get EMBOSS manual here.
Clustal Omega
Best for: Multiple sequence alignments

Clustal Omega in bioinformatics is a next generation sequencing tool designed for doing multi sequence alignments. The genome testing software for Linux supports different input sequence types such as HMM profile and aligning the sequence.
Features of Clustal Omega Alignment Tool:
- Pairwise sequence alignment tools
- Progress alignment for Clustal Omega result interpretation
- Fast guide tree
- HMM profile techniques
- Clustal Omega phylogenetic tree generation
Get Clustal Omega: Available Clustal Omega Download.
BLAST
Best for: Finding match between protein & nucleotide sequences
BLAST or Basic Local Alignment Search Tool helps find similarity amongst biological sequences. The genome testing tool for Linux supports finding match between protein and nucleotide sequences. BLAST in bioinformatics tool also helps with structuring query sequences and mapping such data sets.
Features of BLAST Bioinformatics Tool:
- Managing distance sequences
- Protein-protein comparison and relation
- Sequence aligning using protein constraint
- Searching T cell receptors and immunoglobins
- Finding conserved domains in a sequence
BLAST Bioinformatics Uses:
- Searching protein sequence databases
- Aligning query sequence
- Detecting local aligment sequences
Blast Bioinformatics Tutorial: Get access to BLAST bioinformatics pdf.
Bedtool
Best for: Bedtools genomecov for computing historgrams

Bedtool bioinformatics platform is used for genomic testing and analysis purposes. The application supports different genome formats like VCF, GTF/GFF, BAM and BED. The bioinformatics software for Linux/UNIX and Windows can also be sued for shuffling genomic intervals of different files.
Features of Bedtool bioinformatics Software:
- Targeted DNA capturing
- GC content calculation
- ChromHMM tracks
- RNA-seq coverage analysis.
- DNase hypersensitivity similarity calculation
- Bedtools multicov for reporting alignment counts
Bedtools Download: Bedtools download is available for Windows, Unix and Linux.
Bioclipse Open Source Bioinformatics Tool
Best for: Biological sequencing
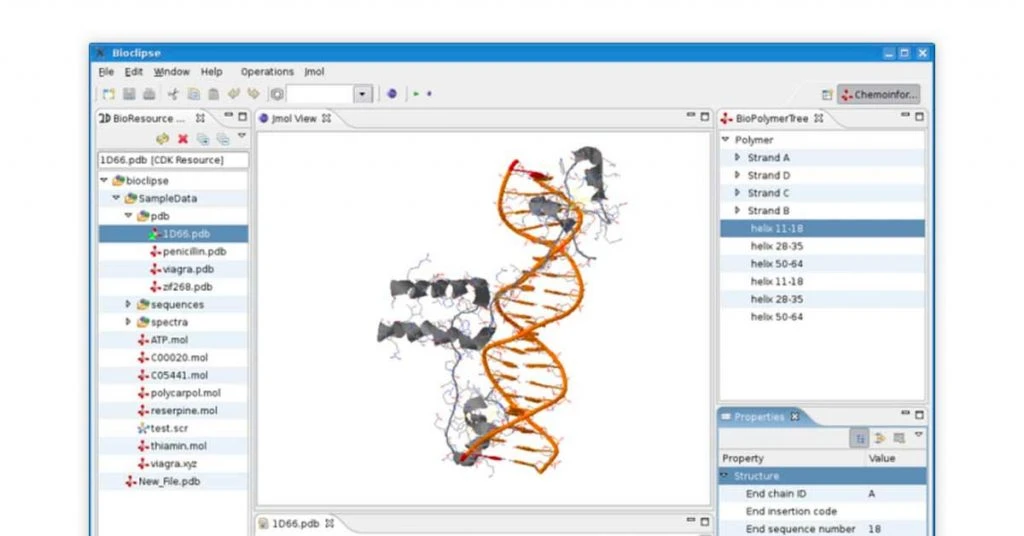
Bioclipse bioinformatics software for Linux is a java-based tool supporting biological sequences like protein, DNA and RNA. The genome sequencing tool is based on plugin architecture for semantic web functioning. Bioclipse is best used for representing molecular structures.
Features of Bioclipse Open Source Bioinformatics Tool
- Single bench for accessing chemo toolkit
- 3D visualization and 2D editing
- Eclipse Rich Client platform
Bioclipse Download: Bioclipse download available for windows and macOS.
Bioconductor
Best for: Real-time biological metadata
Bioconductor is an open source bioinformatics tool that makes use of R programming for analyzing data like oligonucleotide arrays and flow cytometer. You can also deploy these solutions for generating powerful statistical and graphical database.
Bioconductor Bioinformatics Tool Features:
- Digital gene expression
- Real-time biological metadata
- Comprehending high throughput genomic data
- Genomic annotation
- Reproducible research
Bioinformatics Bioconductor Package: Ape package aids, Adegenet, Affy and DEGseq.
Bioconductor Tutorial: Access Bioconductor tutorial.
Conclusion
Sequencing helps understand mutations and variations in human genes to identify disorders and serious diseases like cancers. Bioinformatics databases with their advanced sequencing techniques for genomic testing have proven critical for screening deadly diseases.
FAQs
What is Genomic Testing?
Genomic Testing is a type of test done to study mutations or alterations in genes to identify diseases, food-borne bacteria, and infections.
What is the purpose of bioinformatics tools?
For organizing vast molecular biological data, Developing tools required for analyzing the data and Accurate interpretation of results.
How is bioinformatic tools used in genomics?
Bioinformatics tools help perform genomic tests or next-generation sequencing for studying mutations in genes. These genes help predict the nature of a disease.
You can also check relevant categories:
Somya is one of the most experienced technical writers in the team who seems to be comfortable with all types of business technologies. She is a sensitive writer who ensures that businesses are able to find the right technologies through her writings. She would leave no stones unturned... Read more




























